jejuni invasion in the cells lin ing the gastrointestinal tract. Even though progress has become created in identifying C. jejuni virulence determi nants, the mechanism of cell invasion as well as host cell elements concerned in C. jejuni uptake are less very well defined. Lipid rafts are distinct areas with the plasma membrane that contain high concentrations of cholesterol and gly cosphingolipids. Caveolae really are a distinctive sort of lipid raft. Caveolar membranes have caveolins, which bind cholesterol and type complexes with glycosphingolipids and glycosyl phosphatidyl inositol anchored proteins. 3 members with the caveolin gene family have already been identified. Caveolin one, a 21 to 24 kDa integral membrane protein, is usually a principal part of caveolar membranes along with a key part in the vesicular transport process while in the trans Golgi network.
Caveolin 2 tightly interacts with caveolin 1. Much more specifically, the interaction with caveolin 1 is necessary for transport of caveolin two to your plasma membrane, wherever the 2 proteins kind hetero oligomeric complexes inside of caveolae. Caveolin 2 is really a minor com ponent of the hetero oligomeric complexes, and it is readily degraded during the absence investigate this site of caveolin one. Caveolin 2 has been proposed to act as a co element for caveolae formation, regulating the size and shape in the structures. Pertinent to this study, caveolin 2 is not important for caveolae forma tion, and caveolin 1 and caveolin two are certainly not expressed in all cells. In contrast to caveolin 1 and caveolin two, cave olin 3 is only expressed in striated muscle. Proof from a number of in vitro research has suggested that caveolae play a role in C.
jejuni invasion. Wooldridge et al. demonstrated that read more here therapy of Caco 2 cells together with the polyene antifungal agent filipin III, which binds to and sequesters cholesterol within the membrane, inhibited C. jejuni internalization of human Caco two cells within a dose dependent method. A decade later on Hu et al. per formed similar experiments making use of human INT 407 epithe lial cells, and observed that remedy of those cells with filipin III resulted in a dose dependent reduction in C. jejuni inva sion. Similarly, Watson and Galan found the deal with ment of human T84 cells using the cholesterol depleting compound methyl B cyclodextrin blocked C. jejuni internalization within a dose dependent method. These investigators also reported that transfection of Cos one fibroblast like cells that has a dominant unfavorable mutant of caveolin one, which prevents caveolin one activation by stopping the phosphorylation of tyrosine 14, substantially decreased C. jejuni internalization. To further dissect the importance of caveolae in C. jejuni internalization, the Cos 1 cells have been transfected with a dominant negative type of dynamin II to inhibit caveolae dependent endocytosis.
Monthly Archives: August 2014
By plotting the mean arbitrary fluorescent units from untreated
By plotting the mean arbitrary fluorescent units from untreated and canavanine treated pools, we could clearly determine the can1 can1 deletion strain as extremely enriched inside the population following canavanine treat ment. In the robot assisted experiments, four replicates of a deletant strain for each from the recognized yeast genes encod ing transporter proteins have been spotted onto strong med ium. Growth on a plate containing canavanine identified only the identified canavanine resistant strain can1 can1, in full agreement with published data and with our results from the competi tion experiment described above. We validated the outcomes from both higher throughput experiments by performing development experiments in a BioScreen C instrument, which generates robust development curves under a lot more strictly controlled situations.
We calculated the maximum growth rate from the WT and can1 can1 strains inside the presence of canavanine, and confirmed that, unlike the wild variety, can1 can1 mutants are insensitive to canavanine. In addition, a competition experiment in between canava nine as well as the ON-01910 ic50 native Can1p substrate, arginine, illustrates the fully protective impact of arginine. Both of those benefits suggest that the cellular import of canavanine happens exclusively via Can1p, as reported previously. Drugs with a single protein carrier The two screening procedures identified several transporters which clearly represented the sole transporter responsible for the uptake of a specific drug into yeast cells. The first example is comparable to that of Can1p trans porter and canavanine.
We screened for transporters the full report responsible for the uptake on the anticancer drugs five fluor ocytosine and 5 fluorouracil and, as could happen to be expected, identified that the fcy2 fcy2 mutant was the most resistant strain. Fcy2p is usually a recognized cytosine transporter and is so named due to the fluorocytosine resistant phenotype of its mutant alleles. The analysis of data from pool competitors experiments with diphenyleneiodonium chloride, by plotting the imply arbitrary fluorescence of untreated and treated pools, identified the nrt1 nrt1 deletant as very enriched in the population following DPI treatment. Robot assisted experiments using individual transporter deletants spotted onto agar also identified Nrt1p as the probably DPI transporter. We next performed growth assays on WT and nrt1 nrt1 strains within the presence of increasing DPI concentrations and verified the resistance conferred by the deletion of the candidate transporter. DPI is an inhibitor of reduced nicotinamide adenine dinucleotide phosphate oxidase and related enzymes, and bears some structural similarity to nicotina mide riboside. Furthermore, each nicotinamide riboside and thiamine are known to become transported by Nrt1p.
Because of these final results, one can now derive a z score for
Because of these final results, a single can now derive a z score for every single motif and for that reason rank them in line with their exceptionality. We then worked on modelling the comprehensive distribution of the count of a coloured motif in an ER random graph model. To this objective, we performed a sizable quantity of simulations, working with dierent colour frequencies for the motif and dierent variety of vertices and edges for the graph. We could establish that the Poisson distribution was not proper whereas the Polya Aeppli distribution was a good and greater approximation than the generally utilized Gaussian distribution. The decision of a Polya Aeppli distribution was driven by the following details, motif occurrences overlap in a network, as shown in Figure 1, compound Poisson distributions are specifically adapted to model counts of clumping events, Polya Aeppli approximations are ecient for the count of words in letter sequences.
These final results can in turn be made use of to derive a P value for every single motif, and, thus, to introduce a reduce o for deciding which motifs must be selected for downstream analysis. To our information, there has been no previous function around the signicance of coloured kinase inhibitor P450 Inhibitors motifs in random graphs. This really is the explanation why we started by focusing around the additional common random graph model that may be out there. We are aware that this may not be probably the most suitable model to describe the structure Coloured Random Graph Model. We take into account a random graph G with n vertices V1, Vn. We assume that random edges are independent and distributed as outlined by a Bernoulli distribution with parameter p 0, 1.
Additionally, vertices are randomly and independently coloured as follows. Let C be a nite set of E7080 r dierent colours and f a probability measure on C, f is then the probability for a vertex to be coloured with c C. Inside a metabolic network, the colours of reaction vertices can represent classes of chemical transforma tions, in regulation networks, the colours of gene ver tices can represent functional classes. For dening these classes, the EC quantity hierarchy is classically utilised. Coloured Motif. We consider motifs as introduced in Lacroix et al, a motif m of size k is actually a multiset of k colours m1, mk Ck. Colours from a motif may not be dierent, that’s, a single may well have mi mj for some 1 i, j k. We then denote by sm the multiplicity on the colour c in m. When there is no ambiguity, sm will simply be denoted by Figure 2, Example of a graph and a motif. The motif m happens 3 instances within the graph, at positions s. The notion of multiplicity of a single colour in m are going to be extended to a multiset of colours in Section three. two. Motif Occurrences. We now dene an occurrence of such a coloured motif. To this purpose, we introduce the following notation.
The eluates had been dried, redissolved in 500 uL of mobile phase
The eluates have been dried, redissolved in 500 uL of mobile phase A, filtered and injected in to the HPLC program. Angiotensin peptides we analyzed by reversed phase ODS Aquapore 300 HPLC column, 7 um particle size, applying the gradient five 35% of mobile phase B below a flow of 1. five mL min for 40 min. The angiotensin peptides were identified in line with retention time and peak height of stand ard angiotensin peptides and normalized based on kidney weight. Cell samples and viability assay for flow cytometry Kidney cell enriched fractions in the kidneys with the mouse groups have been ready based on earlier studies. The left kidney was grossly triturated applying sur gical scissors and incubated with an extraction option containing proteinase K and collagenase variety II to dissociate the cells.
The cell extract was filtered through a nylon screen to take away the cell debris, the samples have been washed twice in phosphate buffered saline and stored at 80 C till further evaluation. Cell viability was assessed by propidium iodide selleck ex clusion. A total of 106 cells have been incubated with two uL of PI for 5 min inside the dark at room temperature. Then, cells had been washed with PBS and analyzed using a FACS Canto II flow cytometer. For viability quantification, samples had been acquired in triplicate, and ten,000 events had been utilised for each measurement. Cells have been excited at 488 nm, and PI fluorescence was de tected working with a 585 42 bandpass filter. Information are expressed as the percentage of unstained viable cells. Measurement of intracellular reactive oxygen species The ROS evaluation was performed by flow cytometry as previously described.
P005091 882257-11-6 Dihydroethidium and 2,7 dichlorofluorescein diacetate have been utilized to detect intracellular O2 and H2O2, respectively. Offered its ability to freely permeate cell membranes, DHE has extensively been utilised to monitor O2 produc tion. Upon reaction with O2, DHE is quickly oxi dized to kind ethidium, a red fluorescent product that intercalates DNA and amplifies the red fluorescence sig nal. DCF DA is actually a cell permeant indicator of H2O2 pro duction that is certainly nonfluorescent till oxidation happens within the cell, which converts DCF DA into the fluor escent kind, which remains trapped in the cell. DHE and DCF DA had been added to cell suspensions, which had been then incubated at 37 C for 30 min in the dark, to identify the intracellu lar O2 and H2O2 concentrations, respectively.
Samples that were treated with 10 uM doxorubicin or 50 mM H2O2 for 5 min to create oxidative strain with out cell toxicity, have been made use of as the optimistic handle. Cells incubated with ethanol have been used as the adverse manage. The NO measurements have been performed as previously de 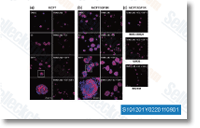 scribed. Briefly, the NO sensitive fluorescent probe four,5 two diacetate was added for the cell suspension, and also the cells have been incubated at 37 C for 180 min in the dark.
scribed. Briefly, the NO sensitive fluorescent probe four,5 two diacetate was added for the cell suspension, and also the cells have been incubated at 37 C for 180 min in the dark.
To elucidate the cis acting elements within the uPA gene promoter
To elucidate the cis acting components in the uPA gene promoter that mediate PB MCM induced uPA transcription, luciferase assays have been carried out by using the p2350 Luc plasmid and several deletion or mutant promoter constructs. In human chondrocytes, the 2,350 30 area of the uPA promoter directed maximal luciferase activity. Sequence deletions from two,350 to 1,872 slightly impaired PB MCM induced uPA promoter activity. Further deletions from 1,872 to 1,700 and mutations in NF B binding sites, nevertheless, lowered PB MCM induced uPA promoter activity by far more than 80% compared with p2350 Luc. We further tested whether or not NF B and AP 1 activations are involved inside the signal transduction path way major to PB MCM induced uPA gene expression.
Human chondrocytes were incubated having a particular inhibitor for NF B or AP 1 for 1 hour, which was followed by stimulation with PB MCM for two hours. The PB MCM induced uPA mRNA expression levels and uPA promoter activity in chondrocytes was drastically lowered natural compound library by means of inhibition with SN50, and partially inhibited with Tanshinone IIA, indicating that NF B is definitely the key transcription element involved inside the regulation of uPA gene induction. To investigate whether NF B binds the uPA promoter area in human chondrocytes, we performed quantitative evaluation of your NF B p65 binding activity in vitro by utilizing TF ELISA kits from Panomics. The remedy of chondrocytes with PB MCM brought on increased NF B p65 DNA binding activity immediately after 0. 5 hours, which remained elevated for a minimum of 1 hour. These benefits have been confirmed by ChIP analysis.
Chromosomal DNA immunoprecipitated using a p65 antibody was sub jected to PCR by utilizing primers made to amplify the uPA promoter region harboring the NF B binding web page. NF B was certainly identified to bind towards the uPA promoter region containing the NF B consensus ATP-competitive MEK inhibitor websites. The JNK and Akt signaling pathways are involved in macrophage induced uPA promoter activity To evaluate no matter whether the inhibition of uPA expression by the JNK and Akt signaling pathways occurs at the tran scriptional level, we studied the effects of specific inhibi tors, siRNA molecules that target JNK, plus a DN Akt on PB MCM induced uPA p2350 Luc promoter and NF B p65 activities. Culturing of your chondrocytes in PB MCM increased the p2350 Luc and NF B p65 activities by five. five and four. 5 fold, respectively, compared with unstimulated cells and just after normalization having a transfection control. Pretreatment from the cells with SP600125 and LY294002, or transfection with JNK siRNA and DN Akt, resulted inside a marked inhibition of both the PB MCM induced uPA promoter activity and NF B p65 activation. Pretreatment with SP600125 and LY294002 brought on a simultaneous and additive inhibition of PB MCM induced p2350 Luc and NF B p65 activities.
We now examined no matter whether distinct classes of BMP respons
We now examined no matter if diverse classes of BMP response may be evoked concomitantly in person dI neurons and irrespective of whether these responses are initiated at dif ferent BMP ligand concentrations. We monitored BMP evoked phosphorylation of Smad1 5 eight as an early step in the classical transcriptional signaling pathway. Smad1 5 8 phosphorylation was measured each by wes tern blot analysis of dI neuronal lysates and by immuno fluorescent labeling of dI neuron cultures. In sister cultures, we also measured development cone collapse, as an instance of an acute response to BMP7, occurring within minutes, and regarded a surrogate for the axo nal orientation response. Development cone collapse in the presence of BMPs was compared by measuring the development cone area in dI neuron cultures, working with ezrin radixin moeisin immunoreactivity to visualize the development cone.
Cultures of dissociated dI neurons had been exposed to BMP7 and BMP6 at two concentrations, 50 ng ml, determined by the observation of dI neuronal specification in explants, and 0.01 ng ml, a concentration sufficient to elicit monocyte chemotaxis. selleckchem At 0. 01 ng ml neither BMP7 nor BMP6 evoked Smad1 5 8 phosphorylation, but at 50 ng ml both ligands stimulated phosphorylation of Smad1 five 8, with phospho Smad1 five 8 labeling detected in 95% of all neurons. In sister cultures, BMP7 elicited similarly robust growth cone collapse at each test concentra tions, causing 46% and 41% decreases inside the average development cone region of dI neu rons. In contrast, BMP6 didn’t elicit growth cone collapse.
While technical issues avoid the use of each ERM and pSmad1 five eight immunoreactivity within the exact same cells, in sister selleck inhibitor cul tures 50% of neurons showed growth cone collapse and 95% showed Smad1 5 8 phosphorylation. These outcomes show that BMP7 stimulates each pSmad1 5 8 activation and development cone collapse in individual neu rons, that BMP6 can elicit only pSmad1 5 8 activation, and that these activities are elicited at distinct thresh old concentrations of BMP7. Sort I BMP receptor signaling participates in inductive specification but not axon orientation Distinct thresholds for BMP evoked inductive specifi cation and axonal orientation raise the possibility that various receptor proteins signal these two activities, supporting the findings suggesting differential roles for variety I and type II receptors in spinal cord and in monocytes. We hence explored no matter whether the inductive and orienting responses of spinal neurons to BMP7 involve the activity of various BMP receptor subunits and or intracellular signaling pathways. Kind I BMP receptors are classically connected with activa tion from the Smad cascade. Nonetheless, knock down experiments have implicated the variety I BMP receptor BMPRIB in roof plate evoked spinal axon orientation.
In general each and every gene is represented by numerous probe s
Normally each and every gene is represented by many probe sets. For every platform we generated the EF statistics for each probe set across the totality of samples. The probe set with all the most robust response across the samples was selected to represent the gene. Explicitly, the probe set using the highest root imply square deviation kind zero was chosen to represent the provided gene. The number of genes defined on each and every plat type had been as follows GPL96 11,807, GPL570 15,983 genes, GPL1261 13,202 genes, GPL85 chip with three,844 genes, GPL1355 chip with 6,341 genes. The database totals 106,101 samples and is searchable on a reasonably quick desktop Pc in 10 minutes per query. Searching the database The query profile is really a statistically thresholded non redun dant list of genes and connected fold values.
Statistical significance is assigned to a fold adjust depending on a sim ple Students t test among various control and treat ment sample expression values. This is when compared with every profile within the database by suggests of a straightforward Pearson regression evaluation, using a correlation coefficient r. The experiments are ranked in line with the buy Neratinib significance. The significance is measured by scaling the correlation towards the typical by a Fisher transformation and measuring the number of standard deviations in the imply. The tion coefficient and N is the quantity of genes making up the correlation. The final ranking score is CMAP combined profiles The CMAP consists of ranked lists of probes for 6,100 separate perturbagen therapies of four diverse human cell lines, with all the ranking determined by response level rela tive to manage.
The treatments are several selelck kinase inhibitor multiples of 1,306 distinctive drug like compounds. To generate responder sets that may be utilized to search SPIED we combined rankings for every separate compound treat ment and converted these into pseudo fold values with linked statistics. The pseudo fold worth is defined by gene and minmax will be the minimalmaximal ranks. Remembering that the highest rank corresponds to the most up regulated gene. The SPIED was searched with CMAP profiles corresponding to folds having a p 0. 05 threshold and with at the very least three replicates. This left 1,218 separate perturbagen probes. We sought to cluster the perturbagens based on predicted target and response profile similarity. The profiles are provided inside the extra file 1 file. Availability of SPIED The SPIED database and related executables are available for download from. The download consists with the SPIED database collectively with executables for browsing SPIED. Supply code files to create the database and carry out query searches are supplied collectively with the executables. Documentation around the database, the execu tables and supply code files can also be incorporated.
Analysis of information All information have been estimated apply
Examination of data All data had been estimated utilizing GraphPad Prism Plan. Quantitative data had been analyzed by a single way ANOVA followed by Tukeys truthfully vital distinction tests amongst individual groups. Information were expressed as suggest SEM. A value of P 0. 05 was deemed major. Effects TGF b1 induces de novo synthesis of MMP 9 and cell migration in RBA one cells To investigate the effects of TGF b1 on MMP 9 expres sion, RBA 1 cells were treated with numerous concentra tions of TGF b1 for that indicated time intervals. The affliction media have been collected and analyzed by gelatin zymography. As proven in Figure 1A, TGF b1 induced MMP 9 expression in a time and concentration depen dent method. There was an apparent up regulation within 16 h and sustained over 24 h.
In contrast, the expression of MMP 2 was buy NVP-AUY922 not significantly modified dur ing incubation with TGF b1. To additional examine whether or not the increase of MMP 9 expression by TGF b1 resulted in the induction of MMP 9 mRNA expression, a RT PCR analysis was performed. The data present that TGF b1 time dependently induced MMP 9 mRNA expression in RBA 1 cells, whereas the expression of a housekeeping gene b actin mRNA was not transformed. There was a substantial improve in MMP 9 mRNA inside four h and sustained over 24 h all through the period of observation. Also, to find out no matter whether the TGF b1 induced MMP 9 expression is dependent on de novo protein synthesis, the cells were exposed to TGF b1 in the absence or presence of actinomycin D or cyclo heximide at a dose known to inhibit transcription or protein synthesis, respectively.
The results display that TGF b1 induced MMP 9 expression was signifi cantly attenuated by pretreatment with both Act.D or CHI in a Omecamtiv mecarbil CK-1827452 concentration dependent method. Additionally, TGF b1 induced MMP 9 mRNA accumulation was atte nuated by pretreatment with Act. D but not with CHI. Additionally, to show the practical activity of MMP 9 expression induced by TGF b1, we evaluated in vitro cell migration of RBA one by a cell migration assay. Immediately after 48 h of TGF b1 incubation, the photographs show that TGF b1 enhanced cell migration was blocked by pretreatment together with the inhibitor of MMP two 9 exercise, suggesting that up regulation of MMP 9 and its activity are essential for improving RBA one cell migration induced by TGF b1.
TGF b1 induces MMP 9 expression and cell migration through a TGF b kind I receptor SB431542, a selective inhibitor of TGF b Sort I recep tor, continues to be shown to abrogate TGF b1 mediated expression of many genes in different cell types. Therefore, we examined irrespective of whether TGF b1 induced MMP 9 expression via TGF bRI, a selective TGF bRI antagonist SB431542 was used for this pur pose. The data reveal that blockade of TGF bRI by SB431542 attenuated each TGF b1 induced MMP 9 protein and mRNA expression. In addition, the involvement of TGF bRI in TGF b1 induced cell migration was characterized by a cell migration assay.
